| Carbon
transformations and ecosystem biogeochemistry |
Soils
harbor the largest pool of carbon in terrestrial ecosystems.
The
carbon found
in soils, referred to as soil organic matter (SOM),
assumes
various compounds with distinct chemical properties, such as
recalcitrance to decomposition and C:N ratio. Old,
recalcitrant
compounds, often with low C:N ratios comprise the majority of SOM, and
are presumed to exhibit the greatest temperature sensitivity of
decomposition. Thus, i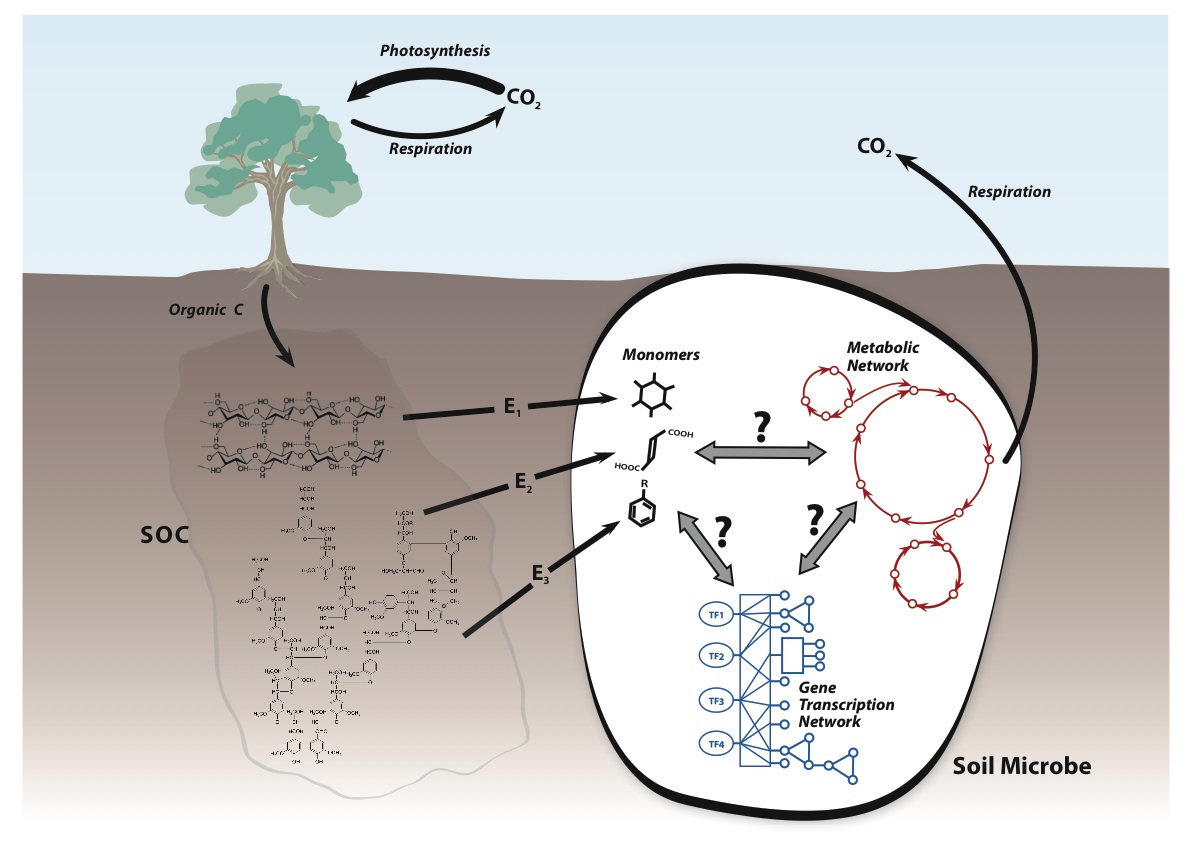 n
order to predict CO2
release from soils in a warmer future, it is necessary to quantify the
temperature sensitivity of microbially mediated SOM decomposition. n
order to predict CO2
release from soils in a warmer future, it is necessary to quantify the
temperature sensitivity of microbially mediated SOM decomposition.
My research employs
theoretical and experimental approaches
to study and quantify the temperature sensitivity of SOM decomposition. This ranges from
characterizing the effects of temperature and edaphic conditions on extracellular enzyme-soil
substrate interactions, to quantifying respiration in the field and in the lab, to analyzing and simulating dynamic models of microbially mediated C transformations. My goal is to link microbial physiology
and community structure to biogeochemical fluxes via equations parameterized with empirical data, which can subsequently be used to project ecosystem response to changing environmental conditions.
|
In aquatic
ecosystems, CO2 is
much less abundant due to its tendency to dissolve and dissociate into
bicarbonate, which poses a challenge for phytoplankton.
However,
increases
in atmospheric CO2 concentrations
will result in increased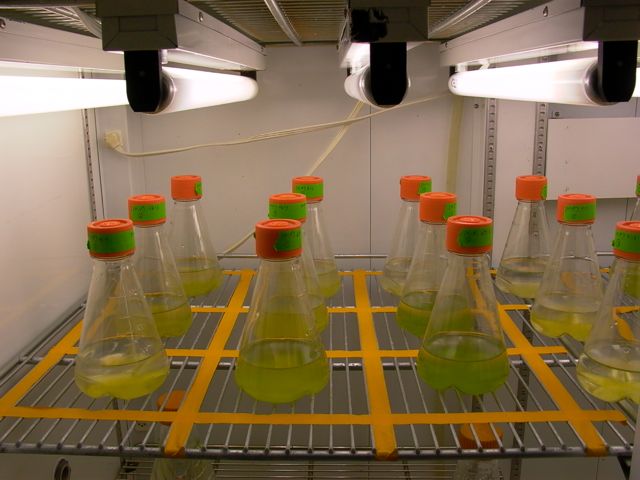  CO2 concentrations
in water, which
increases the
availabiltiy of C for phytoplankton.
Therefore, primary producers in aquatic ecosystems have the
potential to buffer the projected increases in future CO2 concentrations,
if their growth isn't limited by something else. I experimentally
subject phytoplankton and
bacterial
communities to different CO2 concentrations
and N concentrations to determine the potential for aquatic ecosystems
to take up and transform additional C into DOC in response to elevated CO2 concentrations
in the future. CO2 concentrations
in water, which
increases the
availabiltiy of C for phytoplankton.
Therefore, primary producers in aquatic ecosystems have the
potential to buffer the projected increases in future CO2 concentrations,
if their growth isn't limited by something else. I experimentally
subject phytoplankton and
bacterial
communities to different CO2 concentrations
and N concentrations to determine the potential for aquatic ecosystems
to take up and transform additional C into DOC in response to elevated CO2 concentrations
in the future.
|
|

To close the loop on C dynamics in the oceans, I
am working to model DOC transformtions by bacteria using trancriptomic
and FTICR-MS data simultaneously. My goal is develop and
parameterize metabolic models that can be used to simulate DOC dynamics
under a variety of conditions in the oceans.
|
| Ecosystem
stoichiometry and individual nutrient dynamics |
|
Elemental
or stoichiometric ratios of C:N:P in ecosystems are of interest because
they are often remarkably constant across space and through time and
they distill great complexity into a simple metric. Purely physical
forces, for example mixing in the oceans and aoelian deposition on
land, and ecological dynamics, such as photosynthesis,
predation
and evolution, act in concert to determine specific ratio values,
meaning that a given ratio reflects a particular combination of
interacting processes. The challenge is to understand how complex
interactions between abiotic forcing and biological activity conspire
to generate specific ratio values.
Using
mathematical models, empirical ecosystem data, and experimental
phytoplankton cultures, we are studying
how biological processes, 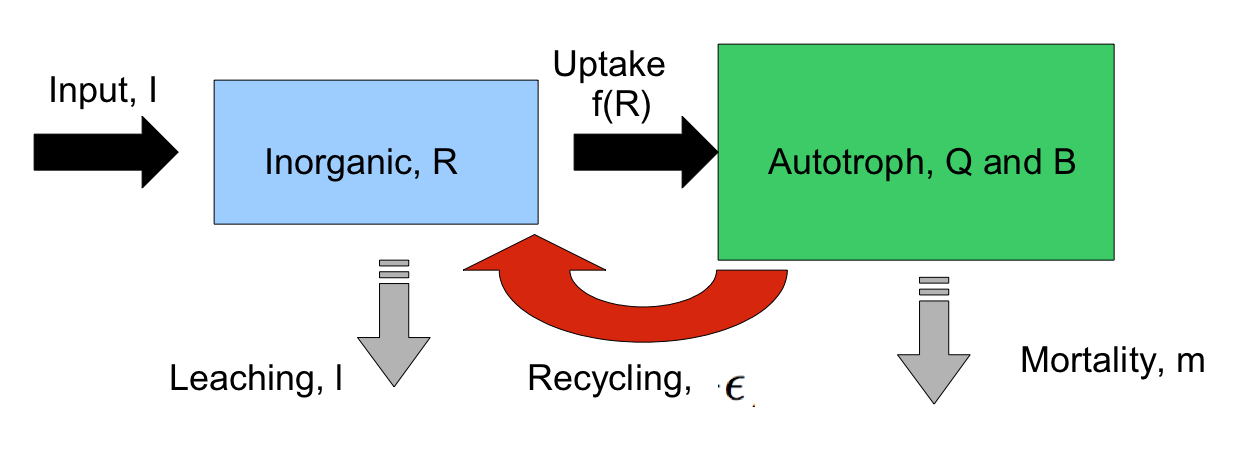 such
as nutrient recycling, affect whole
ecosystem stoichiometry. Models incorporate plant and
phytoplankton
nutrient uptake physiology as well as growth responses to
internal resource
concentrations and relate autotroph nutrient
requirements to N:P stoichioimetry of ecosystem inputs and losses and
recycling. I am developing models that incorporate coupled,
systems-level regulation of inorganic and organic N and P uptake, and
in the lab we are performing experiments to parameterize and test our
models. such
as nutrient recycling, affect whole
ecosystem stoichiometry. Models incorporate plant and
phytoplankton
nutrient uptake physiology as well as growth responses to
internal resource
concentrations and relate autotroph nutrient
requirements to N:P stoichioimetry of ecosystem inputs and losses and
recycling. I am developing models that incorporate coupled,
systems-level regulation of inorganic and organic N and P uptake, and
in the lab we are performing experiments to parameterize and test our
models.
|
|
|
|
|
|
|
|

